Welcome to AmpliSAT (Amplicon Sequencing Analysis Tools)
16/09/2020 Check the new AmpliSAS Github repository.
16/09/2020 Check the new AmpliSAS Docker repository.
14/09/2018 Server was updated and it is running correctly again.
25/06/2018 AmpliCDR3 accepts now as input two paired-end read files and URLs/links to FASTQ files from repositories.
25/06/2018 Improved documentation for AmpliTCR and AmpliCDR3 tools.
04/05/2018 AmpliCDR3 has been optimized for alpha, beta, gamma and delta chains in human and mouse.
AmpliSAT are a set of online tools that make easy the analysis of Amplicon Sequencing experiments.
Amplicon Sequencing (AS) is a useful technique in the genotyping of complex gene families, such as MHC, with multiple loci and copy number variation among individuals (Babik et al. 2010; Lighten et al. 2014).
AS is widely used for taxonomic classification by amplifying a variety of marker genes: small subunit rRNA (16/18S SSU), large subunit rRNA (23/25/28S LSU), internal transcribed spacers (ITS/ITS1,ITS2), cytochrome c oxidase subunit 1 (CO1) or plant specific genes (rbcL, matK and trnH-psbA) (Sogin et al. 2006; Kress et al. 2014; Joly et al. 2014).
Another novel and promising application of AS is the study of antibody and T cell receptor (TCR) repertoires in mammals and birds (Baum et al. 2012; Georgiou et al. 2014; Ruggiero et al. 2015; Robinson 2015).
Click on the desired tool to start the analysis:
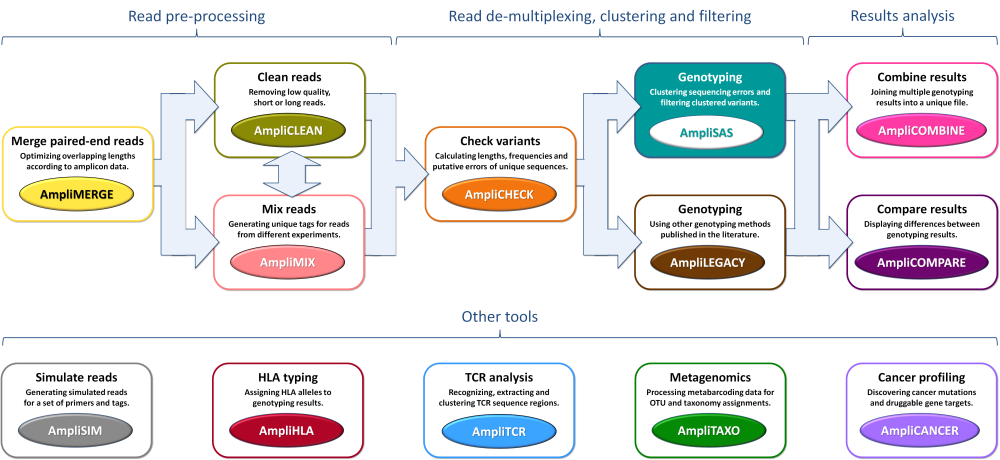
Click to enlarge
Typical amplicon sequencing workflow:
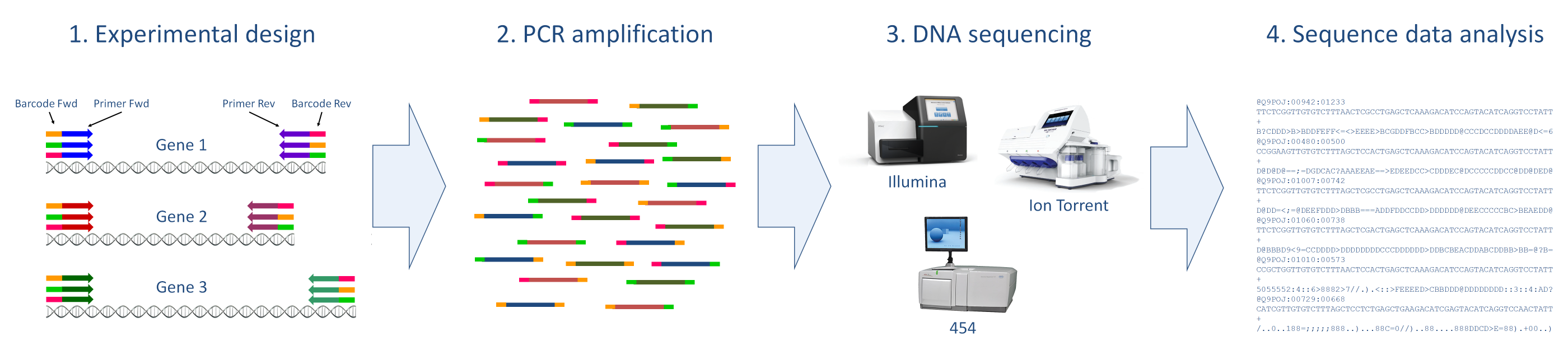
Click to enlarge
Disclaimer
Your use of any of these tools is at your own risk. We do not give any representation or warranty nor assume any liability or responsibility for the data nor the results posted (whether as to their accuracy, completeness, quality or otherwise). Access to these data is available free of charge for ordinary use in the course of research. By visiting the site, you accept our use of cookies and you accept that your data and results will be stored in our server.