AmpliTCR - Amplicon Sequencing T-Cell Receptor segments discovery tool
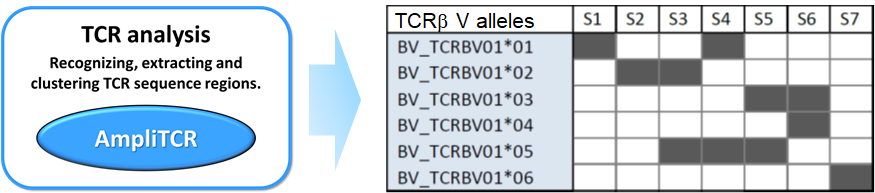
AmpliTCR analyzes a set of genomic or transcriptomic T-cell receptor (TCR) sequences by recognition and extraction of their Variable, Joining, Diversity, CDR3 and/or Constant regions, based solely on pre-defined protein patterns.
AmpliTCR was designed to enable de novo identification of V, J and D segments from NGS targeted/amplicon sequencing reads that cover the entire Variable domain of the receptor (e.g. previously merged 300bp paired-end Illumina reads).
Input DNA sequences are automatically translated into ORFs to extract TCR regions based on previously specified amino acid patterns.
By default, sequences will be clustered with AmpliSAS-like algorithm optimized for Illumina technology and only unique TCR allele sequences segment specific will be retrieved (except for the CDR3 segment).
AmpliTCR requires as input:
- SEQUENCE FILE *: Reads from the experiment in a FASTQ or FASTA format file (compressed or uncompressed).
- TCR REGION PATTERN/S: PROSITE format protein pattern/s to recognise TCR segments as explained in the documentation.
Results can be downloaded on the same page or from an email message after analysis completion.
Results will include a FASTA file containing all the TCR reads matching the desired TCR patterns and FASTA files containing the trimmed sequences of TCR segments according to the patterns.
For more information, read the documentation.
Run AmpliTCR
Disclaimer
Your use of any of these tools is at your own risk. We do not give any representation or warranty nor assume any liability or responsibility for the data nor the results posted (whether as to their accuracy, completeness, quality or otherwise). Access to these data is available free of charge for ordinary use in the course of research. By visiting the site, you accept our use of cookies and you accept that your data and results will be stored in our server.