AmpliTAXO - Amplicon Sequencing TAXOnomy and OTU assignment tool
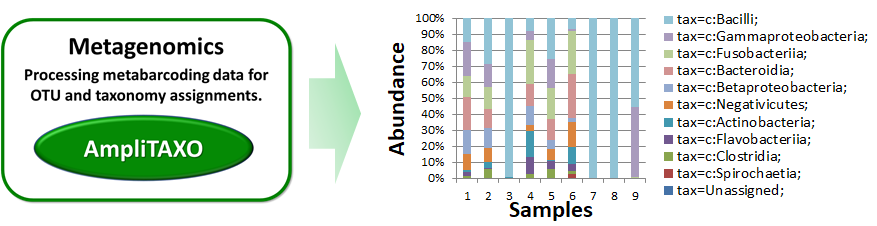
AmpliTAXO analyzes metagenomics data by performing sequence clustering and taxonomic assignments.
Amplicon sequencing data is automatically processed using AmpliSAS algorithm.
Operational Taxonomic Units (OTUs) are retrieved and taxonomic assignments are performed comparing OTUs representative sequences against the most popular metagenomics databases or user provided data.
AmpliTAXO takes as input:
- SEQUENCE FILE: FASTQ or FASTA format file (compressed or uncompressed). Multiple sample/amplicon sequence files should be packed into a unique .ZIP or .TAR.GZ file. Also previously analyzed results can be used as input in AmpliSAS format Excel file.
- AMPLICON DATA 1: primer and tag information in a CSV format file as explained in the documentation.
- REFERENCE DATABASE 2: FASTA file with sequences and headers including taxonomy annotations in UTAX format. For more details read the documentation.
2 Only in case you do not want to use any of the reference databases available below.
Several sets of primers can be used for different regions of the same barcoding gene (e.g. hypervariable nuclear ribosomal RNA regions). In fact it is encouraged to amplify multiple regions to obtain accurate and unambiguous assignments. Other markers can be used if the user provides reference sequences in UTAX format.
An Excel file will be generated with analysis results, two spreadsheets per locus, first one with allele frequencies and second one with expanded redundant allele assignations of high precission genotypes.
Results can be downloaded on the same page or from an email message after analysis completion.
For more information, read the documentation.
Run AmpliTAXO
Disclaimer
Your use of any of these tools is at your own risk. We do not give any representation or warranty nor assume any liability or responsibility for the data nor the results posted (whether as to their accuracy, completeness, quality or otherwise). Access to these data is available free of charge for ordinary use in the course of research. By visiting the site, you accept our use of cookies and you accept that your data and results will be stored in our server.